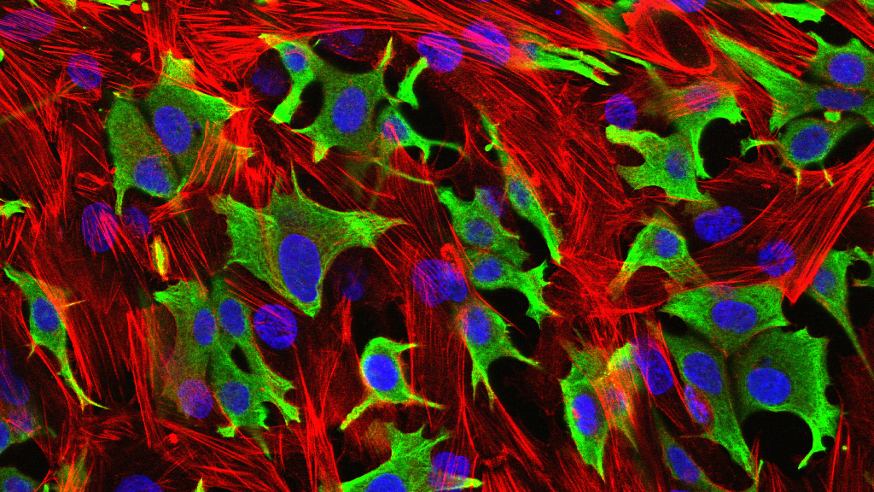
Breast cancer cells (green) invading through a layer of fibroblasts (red). (Luke Henry / the ICR, 2009)
One of the hallmarks of cancer is the uncontrolled growth of cells that leads to tumours forming. Using chemotherapy that disrupts the machinery needed for cell division is commonly used to tackle this.
Some of these drugs target the mitotic spindle — an essential component of cell division — but can cause nasty side-effects and resistance to treatment often develops. So it’s not surprising that efforts are underway to identify new ways to target the mitotic spindle.
Researchers have turned their focus to a group of proteins called kinesins, which play an integral role in maintaining the mitotic spindle, offering an attractive anticancer target.
Professor Ian Collins, Professor of Medicinal Chemistry at The Institute of Cancer Research, London, has recently written a review with Dr Stephanie Myers, an ICR postdoctoral training fellow, on the progress of kinesin inhibitors.
Curious about the role of kinesins in cancer, and how they can be targeted, I caught up with Stephanie to find out more.
The mitotic spindle and centrosomes
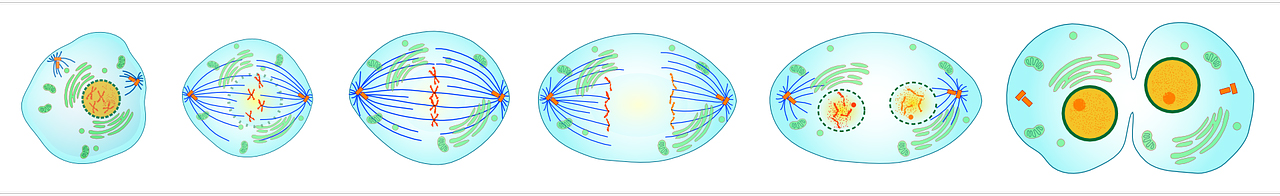
The mitotic spindle consists of a rope of microtubules organised through structures called centrosomes. Normal cells contain two centrosomes, which migrate to opposite poles of the cell during mitosis (cell division). Chromosomes are lined up at the spindle equator to ensure their correct orientation and segregation, and upon mitosis, each daughter cell receives one centrosome and the correct set of chromosomes.
Stages of cell division showing spindle formation, chromosome segregation and generation of two daughter cells [Click to enlarge]
Supernumerary centrosomes (having more than two centrosomes) have been detected in virtually all human cancers. Having multiple centrosomes creates a myriad of problems for the cell, making the genome unstable and consequently leads to programmed cell death. But this raises the question: how do tumour cells survive with supernumerary centrosomes?
The kinesin HSET is believed to play a key role in the survival of cancer cells with multiple centrosomes. By clustering many centrosomes temporarily into the two poles of the cell, the cancer cell is able to disguise itself as ‘normal’ and escape programmed cell death at the mitotic checkpoint. Because healthy cells with the normal two centrosomes are not reliant on HSET for centrosome clustering, HSET offers a tantalising antimitotic target specific for tumour types with a high incidence of centrosome amplification.
The current landscape
So far, two new HSET small molecule inhibitors — AZ82 and CW069 — have been discovered. These inhibitors have been shown to cause centrosome declustering exclusively in cancer cells with amplified centrosomes. Several inhibitors against a kinesin called Eg5 have proceeded further and are in clinical trials, but as a monotherapy, these drugs have been disappointing so far.
Stephanie is working with Professor Collins to develop a new wave of HSET inhibitors. Stephanie tells me that no inhibitor bound crystal structure of HSET is available, so structure-based drug design is a challenge. To add to the hardship, the similarities and differences between the druggable sites on Eg5 and HSET are not yet fully understood.
Fortunately, some clues on how the compounds bind can be gleaned from the structures of other kinesins, and Stephanie remains optimistic in her research. So watch this space for further updates.
Future direction
As is the case with much chemotherapy, the development of resistance to kinesin inhibitors is highly likely, and we have already seen the emergence of resistance to Eg5 inhibitors. Researchers will need to elucidate how resistance to other kinesin inhibitors occurs by looking at the parts of the kinesin that are susceptible to mutation.
Characterisation of where and how inhibitors bind to the kinesins will certainly be essential to the further development of this class of drug.