AI-driven, probabilistic patient motion modelling for radiotherapy planning adaptation
Application closing date: 03/05/26
Project background
Clinical problem
Image-guided radiotherapy (IGRT) offers a transformative opportunity for adaptive cancer treatment by providing high-resolution soft tissue imaging during radiation delivery. However, patient and organ motion continue to present significant challenges that limit treatment precision and accurate reconstruction of delivered doses for quality assurance. Existing motion management strategies are largely reactive, relying on visual assessment, gating, or simplistic surrogate models that do not capture the complex, non-rigid nature of soft tissue deformation. As a result, clinicians must often apply conservative treatment margins, increasing radiation exposure to healthy tissue or risking a miss of the intended target.
Proposed solution
Developing a unique combination of patient motion modelling at The Hawkes Institute at UCL with advanced Bayesian inference with the Computational Imaging group at the ICR, we propose a probabilistic framework to predict and manage soft tissue motion at any time during radiotherapy delivery (see Figure below). This framework will enable prior population-level information, e.g. from AI models trained using a large repository of dynamic cone-beam CT (CBCT), MR-linac (MRL), diagnostic MRI systems, to be combined with pre-treatment imaging data to provide the patient-specific anatomical and motion information. The framework will integrate real-time image or surrogate data acquired during beam delivery, enabling precise and adaptive radiotherapy tailored to individual patients. A key advance of this approach will be full characterization of uncertainty in motion models, including propagation of uncertainty of AI-derived contours of the gross-tumour volume (GTV).
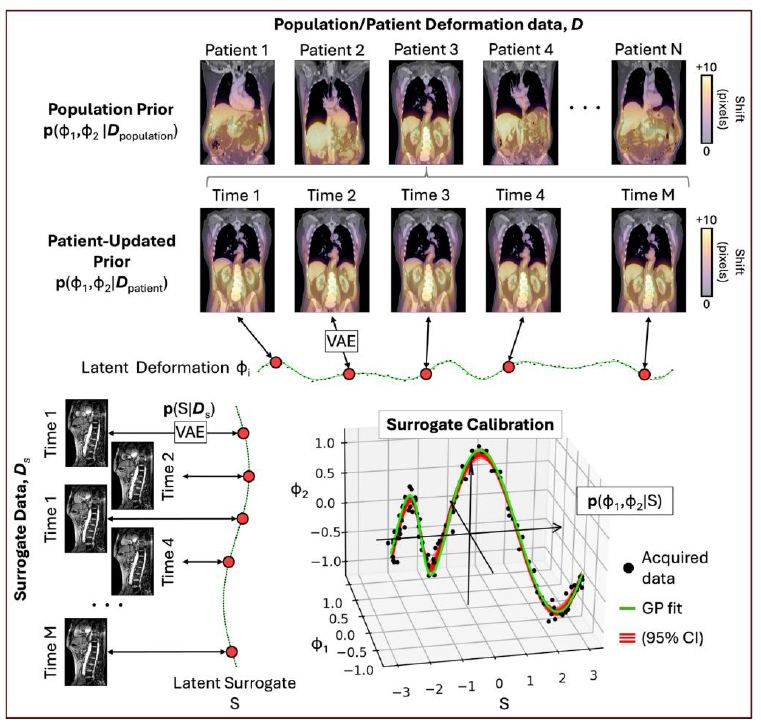
Figure. Overview of the proposed hierarchical Bayesian pipeline for modelling internal anatomical motion during radiotherapy. From existing 4D datasets (top row), the student will construct population-based prior distributions of internal motion. These priors are then refined with patient-specific data, such imaging acquired during previous treatment fractions (second row). Motion is modelled in a low-dimensional latent space of deformations, φ, learned via methods including variational autoencoders (VAE). Finally, a patient-specific probabilistic mapping between internal motion and real-time surrogate signals, S, acquired during treatment (bottom right), enables accurate, real-time prediction of internal anatomy. This approach enhances understanding of motion-induced dose variation and supports potential real-time treatment adaptation. Note: the notation p(a, b | c, d) denotes the probability distribution over unknown parameters a and b given observed data c and d. CI = confidence interval.
Project aims
1. Develop Bayesian models using VAEs to learn optimized patient-specific internal motion priors from high-dimensional 4D imaging deformation fields.
2. Refine and update individualized motion priors over treatment using additional 4D imaging from previous radiotherapy fractions.
3. Calibrate probabilistic mappings from external surrogate signals to latent internal motion for real-time organ and target position estimation.
4. Integrate Bayesian forecasting and uncertainty calibration to predict future motion with low latency and validated probabilistic accuracy.
Further details & requirements
Year 1: Development of Foundational Bayesian Motion Models
During the first year, the student will focus on establishing robust patient-specific models of internal anatomical motion. Building on existing methodology, they will develop Bayesian frameworks, including methods such as deep conditional variational autoencoders, to learn prior distributions of internal motion from high-dimensional 4D deformation fields. Emphasis will be placed on determining the optimal number of latent variables required to balance dimensionality reduction with anatomical fidelity. They will also explore conditioning these models on baseline patient images to improve individual specificity. By the end of Year 1, they are expected to have a validated pipeline capable of generating reliable motion priors for abdominal radiotherapy patients, both for fast moving anatomy (e.g. pancreas) and slower moving anatomy (e.g. bladder).
Year 2: Personalisation and Surrogate Calibration for Real-Time Estimation
In Year 2, the student will refine and personalise these priors by incorporating additional 4D data from previous treatment fractions, allowing continuous adaptation throughout the radiotherapy pathway. Parallel to this, they will develop and calibrate probabilistic mappings that link external surrogate signals, including multiplanar MRI, surface imaging, or physiological monitors, to the latent internal motion space. They will compare established methods (e.g., Gaussian Processes) with modern deep-learning models including transformers, determining which approaches best capture complex nonlinear relationships. These calibrated models will provide real-time posterior estimates of organ or target motion during treatment.
Year 3: Forecasting, Integration, and Comprehensive Evaluation
The student will focus on integrating Bayesian forecasting techniques to predict short-term future motion with minimal latency, essential for real-time radiotherapy adaptation. These methods will include GP-based temporal forecasting and physiology-informed Bayesian models capable of anticipating anatomical changes such as bladder filling or respiratory drift. Rigorous evaluation will be performed using digital phantom simulations and post-treatment MRI studies on the MRL and Radixact systems. We will assess both predictive accuracy and uncertainty calibration to ensure reliable, clinically actionable probabilistic outputs.
Year 4: Manuscript and thesis writing
The final year will be spent developing the material into publishable content and writing the thesis for final submission.
| Pre-requisite qualifications of applicants: | First or 2:1 BSc (or MSc) in Physics, Computer Science, Engineering, Mathematics or a closely related quantitative discipline. Desirable: experience with medical image analysis, Python, and basic statistics. |
| Intended learning outcomes: | · Artificial Intelligence and Machine Learning as applied to healthcare, including exposure to all relevant software tools (PyTorch, Monai, TensorFlow, Data Version Control [DVC], version control), bespoke deep-learning model design and robust model training/validation. · Practical experience in cancer research, including the development and conduct of clinical patient trials and data management. · Practical experience with Magnetic Resonance Imaging including development of imaging protocols and data analysis, both through phantom and patient scans. · Fundamentals of radiotherapy, including the patient workflow, with an understanding of how different imaging modalities are used to optimise patient positioning and safety considerations · Communicate research findings to both scientific and clinical audiences through data visualisation, high-impact manuscripts, and conference presentations. · Collaborate effectively across disciplines, critically appraising literature and emerging technologies in imaging, oncology, probability theory, radiotherapy, and biomarker discovery, while adhering to regulatory requirements for software/AI development |
2. McClelland, Jamie R., et al. "A generalized framework unifying image registration and respiratory motion models and incorporating image reconstruction, for partial image data or full images." Physics in Medicine & Biology 62.11 (2017): 4273.
3. Kingma, Diederik P., and Max Welling. "An introduction to variational autoencoders." Foundations and Trends® in Machine Learning 12.4 (2019): 307-392.
4. Kalantar, Reza, et al. "Domain-Adaptive and Per-Fraction Guided Deep Learning Framework for Magnetic Resonance Imaging-Based Segmentation of Organs at Risk in Gynecologic Cancers." Advances in Radiation Oncology 10.4 (2025): 101745.
5. Shojaei, Mehdi, et al. "A robust auto-contouring and data augmentation pipeline for adaptive MRI-guided radiotherapy of pancreatic cancer with a limited dataset." Physics in Medicine & Biology 70.3 (2025): 035015.
6. Huang, Yuliang, et al. "Surrogate-driven respiratory motion model for projection-resolved motion estimation and motion compensated cone-beam CT reconstruction from unsorted projection data." Physics in Medicine & Biology 69.2 (2024): 025020.
7. Huttinga, Niek RF, et al. "Gaussian Processes for real-time 3D motion and uncertainty estimation during MR-guided radiotherapy." Medical Image Analysis 88 (2023): 102843.
8. Rasmussen, Carl Edward. "Gaussian processes in machine learning." Summer school on machine learning. Berlin, Heidelberg: Springer Berlin Heidelberg, 2003. 63-71.
9. Vaswani, Ashish, et al. "Attention is all you need." Advances in neural information processing systems 30 (2017).
10. Segars, W. Paul, et al. "Application of the 4-D XCAT phantoms in biomedical imaging and beyond." IEEE transactions on medical imaging 37.3 (2017): 680-692.