-embed---874x492.jpg)
Test tubes. Credit: Jan Chlebik for the ICR, 2011.
I have written before on this blog about small-molecule chemical probes – emphasising the need for high-quality tools that can be used with confidence to understand the function of proteins of interest and to help validate them (or invalidate them) as potential drug targets.
Plenty is happening in the world of chemical probes, and in this post I will be reporting back on recent discussions at the American Association for Cancer Research (AACR) annual meeting in Chicago, and reflecting on an exciting new initiative involving industry donating chemical probes and associated data to the biological research community. But first, some wisdom from Indiana Jones.
Choosing wisely
In the 1989 Steven Spielberg movie Indiana Jones and the Last Crusade, the archaeologist and adventurer Indiana Jones (played by Harrison Ford) is on a quest to rescue his father (Sean Connery) and find the Holy Grail before the bad guys beat him to it.
There is a crucial scene, set in the depths of the cliffside Grail Temple that is defended by the Grail Knight. Here, the baddie collaborator Walter Donovan tries to select – from the many that are on display – the particular goblet that is the True Grail which will offer the drinker eternal life.
Donovan, who has previously shot Indy’s father, surveys the collection and asks the Grail Knight: ‘Which one is it?’ The Grail Knight replies: ‘You must choose. But choose wisely, as the True Grail will bring you life, and the False Grail will take it from you.’ Tough choice.
Donovan drinks from a gaudy, jewel-studded gold chalice selected with the help of the duplicitous, femme fatale archaeologist Dr Elsa. Upon drinking, Donovan ages horribly and shrivels away to dust in horrifying fashion – because although flashy his chosen goblet was a False Grail.
 The Grail Knight confirms the obvious: ‘He chose… poorly.’ Quite an understatement.
The Grail Knight confirms the obvious: ‘He chose… poorly.’ Quite an understatement.
Indy of course chooses wisely, selecting the True Grail – an honest, fit-for-purpose wooden cup. He drinks from it without ill effect and the Grail Knight informs him: ‘You have chosen wisely.’ Indy uses the True Grail cup to save his dying father, and all ends happily ever after.
Image: Harrison Ford as Indiana Jones. Source: WikiCommons. Licensed under Creative Commons (CC BY-SA 2.0)
Have a look here if you want to be convinced of the importance of choosing wisely and the appalling consequences of choosing poorly.
You can see where my cinematic metaphor is going. The superficially attractive False Grail chalice represents the poor-quality chemical probe, lacking the required solid properties despite the jewelled attractions of high historic citation rates and glitzy (but inaccurate) descriptions in vendor catalogues. Choosing such a flawed probe can only end in disaster.
The sturdy wooden True Grail goblet, unflashy but skilfully and effectively constructed, is the high-quality fit-for-purpose chemical probe. It possesses all the essential properties, and selecting it for your work gives the true and correct result – even if not eternal life.
Chemical probes at AACR
In my previous post on this blog I wrote about how expert probe developers must ensure that the key properties or ‘fitness factors’ of a chemical probe – such as potency, selectivity and cell permeability – are of the required quality level.
And I have pointed out the importance of getting the message out to the basic biological and biomedical research community that making the right choice of chemical probe is absolutely essential if the biological results obtained are to be trusted.
It was with the latter point in mind that I used the Indiana Jones metaphor at a session I chaired at last week’s meeting of the American Association of Cancer Research.
The session was entitled: ‘The use and abuse of chemical probes – ensuring best practice for interrogating biology and target validation’, and the aim was to promulgate the message of choosing chemical probes wisely that Professor Julian Blagg and I wrote written about in a Perspective article in Cancer Cell last year.
I introduced the session by defining a chemical probe as a small-molecule compound aimed at interrogating the function of a specific protein in cells or organisms.
I emphasised the use of the Pharmacologic Audit Trial concept that my colleagues and I initially developed for use in mechanism-based drug discovery and development – where evidence is obtained that the proposed probe has defined chemical structure and purity, binds to the target of interest and ideally in a way that is understood structurally, shows evidence of target engagement and functional modulation in the cell or organism, and elicits a phenotype associated with the effects on the target.
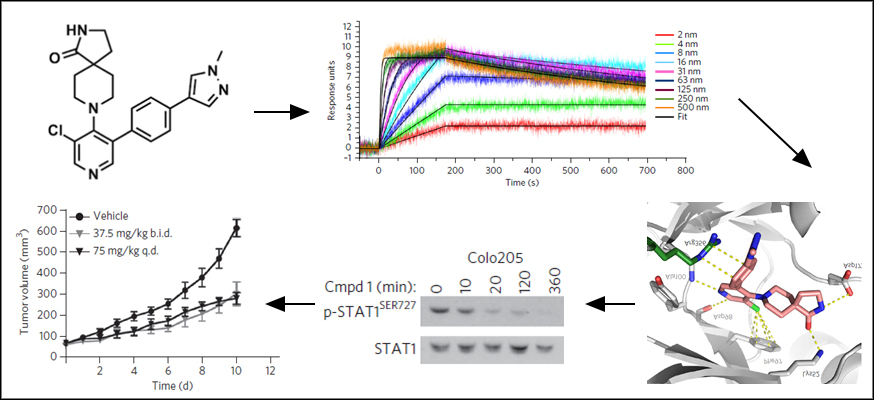
Image: Pharmacological Audit Trail for a chemical probe, in this case a CDK8/19 inhibitor
The aims of the session were to:
- Become familiar with the properties of high-quality chemical probes
- Recognise examples of the good, the bad and the truly ugly
- Appreciate current challenges with the selection and use of chemical probes
- Understand how we can be more discriminating
- Encourage best practice in the field.
I stressed that although chemical probes could be purchased from vendors for around $150, the costs of using poor-quality tools runs into billions of dollars.
I gave examples of continued use of no-longer-suitable chemical probes and examples from our own work showing the value of high-quality inhibitors of CDK8/19 and of pirin. I also illustrated our design of a proteolysis-targeting chimera (PROTAC) for pirin and how this can be used to show in-cell target engagement of chemical probes.
I strongly emphasised the value of the Chemical Probes Portal as an expert-curated ‘go to’ web resource to help researchers choose the best available chemical probe.
Speaking next, my colleague Albert Antolin here at The Institute of Cancer Research, London, explained the synergistic value of our Probe Miner resource, which builds on our integrated Big Data drug discovery and translational knowledgebase canSAR – of which version 4 launched in March this year.
I have previously discussed the advantages of our Probe Miner resource which provides objective assessment of chemical probes based on the integrated analysis of more than 1.8 million compounds against 2,220 human targets.
Albert talked about the new updated version of Probe Miner which has just been released, fully updated to include data from canSAR version 4 and ChEMBL_23. Albert showed examples to emphasise the complementary nature of Probe Miner and the Chemical Probes Portal and plans to integrate these synergistic resources more closely.
Next in the session, Milka Kostic spoke from her experience as a former editor of Cell Chemical Biology and now as Programme Director in Chemical Biology at the Dana-Farber Cancer Institute. Milka talked about the chemical probes ecosystem and argued that we are all responsible for improving the quality of chemical probes including probe developers, users, vendors, funders and journals.
Milka emphasised that: “With the exception of the domain-specific journals, most journals in life sciences don’t have specific standards for validating and reporting chemical probes.” She argued that improving probe standards in non-specialist biology journals would make a real difference.
Milka also set out five components of a push-pull strategy to improve the use of chemical probes, involving:
- Community
- Infrastructure
- Funding
- Outreach
- Updated modus operandi.
Expanding on the last bullet point on ways of working, Milka challenged the community to follow the example of the rigorous standards set, and impact achieved, by the Protein Data Bank (PDB) and also highlighted the non-profit plasmid repository addgene.
Concluding the session, Stephen Frye from UNC Eshelman School of Pharmacy opened his talk by making a key point that cannot be emphasised enough: chemical probes and drugs are different.
Drugs are medicines that need to be safe and effective, but they do not necessarily need to be selective and in many cases hitting multiple targets is beneficial. In contrast, probes are designed to ask a specific question about biology and need to be as selective for the desired target as possible.
Using examples from his own work, Stephen illustrated many important points about chemical probes in the areas of methyl lysine readers; chromodomain cancer relevance; chromodomain and polycomb complexes involved in transcriptional repression in development; and progress towards an in vivo probe for PRC1 chromodomains.
Stephen also showed illuminating examples of how publication of chemical probes from his group has led to productive collaborations on chromodomains which in turn have resulted in new biological knowledge and potential therapeutic applications emerging.
A subsequent session organised under the auspices of the AACR Chemistry in Cancer Working Group and chaired by Angela Koehler (MIT) showed many great examples of the power of chemical biology and chemical probes, as presented by Angela herself, Sarah Buhrlage (Dana Farber) and Dan Nomura (Berkeley), including cutting-edge approaches to ‘drugging the (previously) undruggable’.
Visit the chemical probes web resource – Probe Miner – released to the research community and led by Dr Bissan Al-Lazikani, Professor Paul Workman, Dr Albert Antolin and colleagues.
Donated chemical probes for open science
There are many exciting new initiatives in the field of chemical probes and one of the impressive features has been the amount of pre-competitive, open innovation model activity.
Despite this progress, a limitation has been that although the pharmaceutical industry has generated numerous high-quality chemical probes, some of which have been made available to academia, the detailed probe-associated data and inactive control compounds are generally not accessible.
A new article just published in eLife (on which Julian Blagg is co-author) describes the initial results from a consortium of seven pharma companies, facilitated by the Structural Genomics Consortium (SGC). This group has released detailed information on 70 high-quality chemical probes to enable investigations by the research community.
In addition to the publication, the information, including recommendations on usage, is freely available via a website.
The peer-reviewed probes meet the following criteria:
- Potency <100nM (IC50 or KD)
- Selectivity within a target family >30-fold
- Extensive off-target profiling outside the target family
- 100x less potent control compound available
In addition to the obvious benefit of extending the number of protein targets for which high-quality chemical probes are available, I am particularly pleased to see that inactive control compounds will be systematically sought and provided.
Although it is common to talk about a target ‘probe set’ involving active compounds from at least two chemical series and inactive control compounds, reagents to enable this are not routinely accessible.
Chemically distinct probes for the same target frequently (but not always) have different off-targets and therefore the parallel use of two of these can help reduce the risk that an observed effect of a chemical tool is not in fact mediated through the target of interest.
In the case of inactive control compounds, testing of these should lead to loss of the various effects seen with the paired active probes and if this is not the case then off-target effects can be suspected. Overall, careful application of a suitable quality ‘probe set’ can build confidence (or not) that chemical probes are acting on the intended target – much more than would be the case with a single probe.
Also of note is that: ‘All approved probes are measured against the same quality criteria and will be profiled in assay panels comprising more than 500 assays, including broad panels of pharmacologically active targets such as GPCRs, kinases, ion channels and proteases to identify off-target activities.’
This is important because the ability to have proposed chemical probe compounds tested across large numbers of targets is commonly not available in academia and is a major limitation in the comparative assessment of chemical probes.
Compounds are made available through an online system without complicated contractual arrangements under a simple web-accessible Open Science Trust Agreement.
Together with other resources discussed in this blog, the Open Science Probes resource represents a valuable new initiative. Ensuring that all of the resources are interlinked will make them even more useful across the life-science community.
The benefits will be faster progress in the use of chemical probes to interrogate biological and disease mechanisms, more robust results, better target validation and accelerated therapeutic innovation. Ultimately, more patients will benefit.