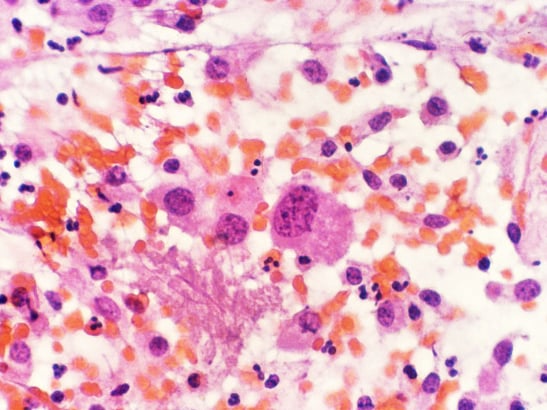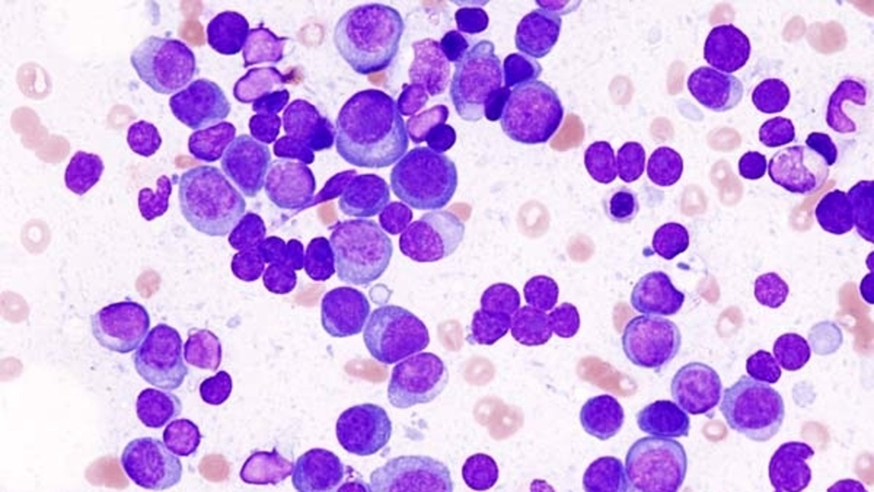
A histopathological image of multiple myeloma. Image by KGH. CC license BY-SA 3.0)
From our eye colour to our height, all of us display great variety on a physical level. We display a similar variety on a genetic level too. Variation in our DNA, accumulated throughout our lifetime, is termed somatic mutations. Depending on where these mutations are, they can be harmful, with some leading to cancer.
Multiple myeloma is a cancer characterised by uncontrolled growth of plasma cells in bone marrow. Myeloma remains incurable and its complex biology is not properly understood. But by looking at somatic variations in DNA, we can build a picture of the landscape of mutations which occur in myeloma.
Work in Professor Richard Houlston’s lab at The Institute of Cancer Research, London, aims to shed new light on the biology of myeloma by piecing together how these mutations fit into cellular pathways.
Byte-sizing Big Data
Previously, most studies have only focused on studying somatic mutations in the 2% of the genome responsible for writing the instructions for protein – the coding regions.
The presence of variants inside a gene can act like a typo in a set of genetic instructions, and can cause problems in how the instructions are read and understood by the cell. This, in turn, can lead to a dysfunctional protein.
However, if we only consider these coding regions, there is 98% of the genome which remains unexplored. Fortunately, advances in technology have driven down the cost of DNA sequencing and it is now feasible and affordable to sequence whole genomes.
In the case of myeloma, a large collaborative project by the Multiple Myeloma Research Foundation called CoMMpass has now sequenced the genomes of 765 patients.
To give you an idea of how large this dataset is, imagine one byte (a unit of data) is a single grain of rice, and a page of writing (3,000 bytes) is equivalent to three cups of rice. The size of this dataset was 10 terabytes (10×1013 bytes) which is equivalent to 20 container ships of rice.
Here, in an abundance of data, Phuc Hoang’s PhD project embarked. His first task was to bite-size the vast amount of data into something the ICR’s supercomputer could handle.
We are training the next generation of researchers; by supporting a PhD studentship, you could help us advance personalised prostate cancer treatment.
Piecing together pathways in myeloma
Phuc successfully narrowed down his search of the genome by employing a strategy which looked at the regions implicated in regulating the activity of our genes.
These regions can be defined by marks which delineate the DNA as ‘regulatory’ and mutations in these regions have the potential to change the activity of genes, like using a dimmer switch to turn a light bulb up and down.
In the paper, Phuc explored which genes and their regulatory regions were mutated more frequently in myeloma patients, finding 40 genes which were significantly somatically mutated.
Of these 29 had been previously implicated in myeloma, while 11 represented novel findings. He found that the regulatory region of the PAX5 gene was recurrently mutated. This gene has an important role in the development of B-cells, an early precursor of the plasma cell.
The next step was to piece these individual genes into pathways which may be affected in myeloma. Some of the pathways identified included those related to DNA damage and cell death, which are intrinsic to cancer, as well as pathways relating to blood cell development and MAPK/NF-kB signalling, which are central to myeloma.
Signatures on the myeloma landscape
Mutational processes can leave characteristic footprints on the genomic landscape and these are termed signatures. The presence of these signatures was investigated in the myeloma dataset and of particular interest, a signature which is attributed to a gene called AICDA was found.
There was evidence that this gene, which causes single-letter changes in the DNA code, acts at an early stage in the development of myeloma.
This gene has previously been associated with rearrangements of DNA called translocations in B-cells. A translocation occurs when a whole section of DNA is shuffled to a different location and these are a hallmark of approximately 50% of myeloma cases.
The study suggested that the pattern of mutations caused by AICDA was more widespread than previously thought. And it found evidence that genes, including an important gene called TP53 which in its intact form helps protect against cancer, were targeted by an enzyme called APOBEC.
This enzyme is responsible for a DNA signature which has previously been associated with poor prognosis in myeloma patients.
Overall the study has helped build a picture of the mutational landscape of myeloma. The research can provide a foundation on which to investigate the biology of the disease, offering the opportunity to identify potential drug targets and the prospect of future personalised therapy in myeloma.
The next step will be to take candidates from the analysis and study them in cell-based experiments in the laboratory. It will be exciting to see what results this biological validation provides.